Towards accurate, contiguous and complete alignment-based polyploid phasing algorithms.
Omar Abou Saada, Anne Friedrich, Joseph Schacherer.
Genomics. 2022. doi:10.1016/j.ygeno.2022.110369
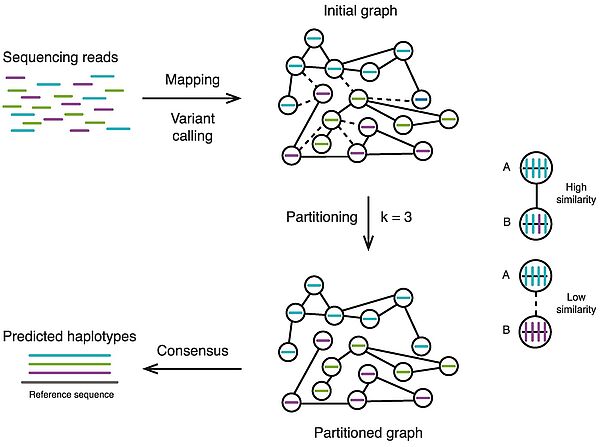
Phasing, and in particular polyploid phasing, have been challenging problems held back by the limited read length of high-throughput short read sequencing methods which can't overcome the distance between heterozygous sites and labor high cost of alternative methods such as the physical separation of chromosomes for example. Recently developed single molecule long-read sequencing methods provide much longer reads which overcome this previous limitation. Here we review the alignment-based methods of polyploid phasing that rely on four main strategies: population inference methods, which leverage the genetic information of several individuals to phase a sample; objective function minimization methods, which minimize a function such as the Minimum Error Correction (MEC); graph partitioning methods, which represent the read data as a graph and split it into k haplotype subgraphs; cluster building methods, which iteratively grow clusters of similar reads into a final set of clusters that represent the haplotypes. We discuss the advantages and limitations of these methods and the metrics used to assess their performance, proposing that accuracy and contiguity are the most meaningful metrics. Finally, we propose the field of alignment-based polyploid phasing would greatly benefit from the use of a well-designed benchmarking dataset with appropriate evaluation metrics. We consider that there are still significant improvements which can be achieved to obtain more accurate and contiguous polyploid phasing results which reflect the complexity of polyploid genome architectures.